Spatial Profiling using the GeoMx RNA Assay for NGS Readout: Robust Technology for Identification of Novel Biomarkers and Early Detection of Melanoma
Early diagnosis of melanoma is critical for improved clinical outcomes and survival of patients. Yet disagreement in traditional histology-based diagnosis frequently occurs. Despite currently available molecular technologies, melanomas characterized by low cellularity and heterogeneity can hinder accurate diagnosis.
To address these challenges, Dr. Maija Kiuru, Director of Molecular Dermatopathology at the University of California, Davis, used the NanoString GeoMx™ Digital Spatial Profiler (DSP) to spatially resolve specific cell types within skin tissue. GeoMx DSP coupled to next generation sequencing (NGS) readout enabled Dr. Kiuru to profile RNA expression of 1,412 genes in formalin-fixed paraffin-embedded (FFPE) skin biopsies to identify novel biomarker candidates for melanoma.
During the recent webinar, Cell Biology 2019, Dr. Margaret Hoang, lead Scientist on the GeoMx NGS Assay development at NanoString, explained: 1) how the GeoMx DSP technology enables spatially resolved mRNA and protein analysis, and 2) how users can achieve reliable, reproducible analyses of FFPE tissue samples. The use of the GeoMx RNA Assay with NGS readout allows users to comprehensively profile the cancer transcriptome. She concluded her presentation by stating that, in addition to capturing tissue morphology in a spatial molecular fingerprint, this technology also has a potential to detect splice variants in cancer tissues.
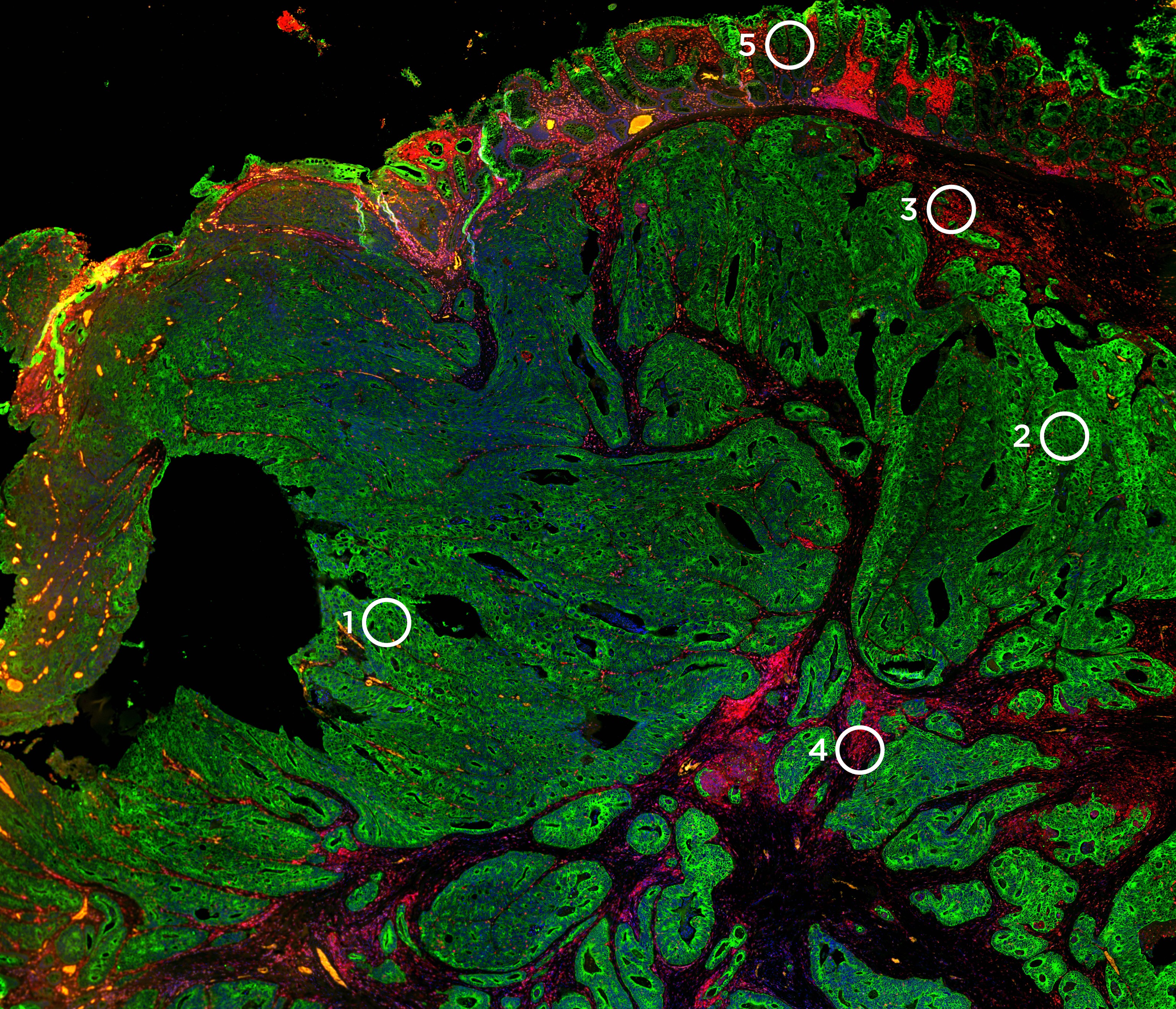
Next, Dr. Kiuru presented data from her team’s study “Identification of cell type-specific RNA biomarker candidates in melanocytic tumors using digital spatial profiling.” Her team evaluated technical performance and reproducibility of the assay on four case types: common nevus, dysplastic nevus, melanoma in situ, and melanoma. Regions of interest (ROI) of 200µm circles in FFPE sections were selected covering five cell types categories: immune-rich, Keratinocytes/melanocytes, mixed, melanocyte-rich, and control epidermis. NGS readout from DSP tags performed well, with high reproducibility between replicate libraries and between serial sections. Moreover, her team validated the melanocyte-associated gene counts and immune gene counts with immunofluorescence staining analyses.
After the technical evaluation, her team moved on to “Study Phase II: compare gene expression across tumor types & cell types”. In Phase II, three patients per case type (12 total) were analyzed. ROI clustering suggested that each case type may have a distinct molecular signature. They further compared differential expression of known melanomagenesis-associated genes and PRAME (PReferentially expressed Antigen in MElanoma) between benign melanocytic nevi vs. melanomas. PRAME was overexpressed in melanoma specifically in melanocytes. Moreover, antigen presentation genes were elevated in the melanoma tumor microenvironment. Overall, Dr. Kiuru’s study verified the utility of GeoMx DSP on FFPE tissues from melanocytic tumors.

