How to Resolve the Complexity of Autoimmune Diseases
What are Autoimmune Diseases?
Autoimmune diseases are chronic inflammatory conditions, life-long debilitating illnesses with increased mortality, and high healthcare costs for therapy and care. They are systemic conditions characterized by activation of either the adaptive immune system—rheumatoid arthritis, lupus, multiple sclerosis— or the innate immune system—Crohn’s disease, ulcerative colitis. Other diseases such as psoriasis or ankylosis spondylitis exhibit activation of both immune responses.
What are the Challenges in Treating Autoimmune Diseases?
Autoimmune pathogenesis is challenging because it is multifactorial, deriving from a complex interaction between genetic and environmental factors. Such complexity makes it unlikely to find single biomarkers that reveal both the driver of the disease and identify an effective target for treatment. It is more likely that panels of biomarkers evaluated at different stages of progression will be used for disease detection and to guide treatment decisions.
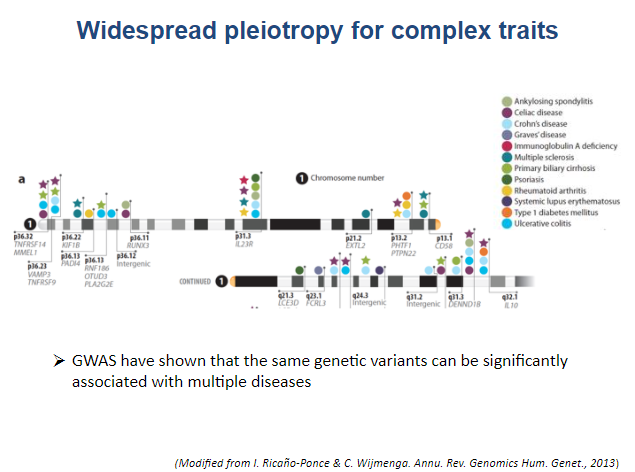
With the development of high throughput sequencing technologies, genome-wide association studies (GWAS) have uncovered hundreds of genetic variants that influence the risk for chronic inflammatory diseases. These studies have uncovered overlapping loci that lead to a predisposition for several diseases, meaning that some chronic inflammatory illnesses share disease-associated risk variants. Yet functional studies aimed at delineating the causal genetic variants and biological mechanisms underlying the observed associations with disease risk are lagging.
There are many factors causing this discrepancy. The complexity and interindividual variability of the immune system and the inflammatory pathways shared by different autoimmune diseases are major challenges to understanding the biological mechanisms behind the disease. The limited amount of material available—blood, lysates, FFPE, circulating RNAs—is also an important consideration.
What is the Solution?
NanoString has developed curated, ready-to-use gene expression panels to measure various aspects of the autoimmune response. Like any other nCounter panel, they are easily customizable to address specific research questions and can be used on small specimens.
In two of our on-demand webinars, former NanoString scientist Kirsty Maclean, Ph.D., and NanoString customer Lars Rogge, Ph.D. from the Institut Pasteur guide us through the challenges that scientists have encountered in defining the functions of genes associated with autoimmune diseases and the solutions they adopted.
In the first webinar, Dr. Mclean explains how nCounter technology can support scientists in their quest for biomarkers and therapeutic targets in autoimmune diseases. We learn how Joan Wither and colleagues at the University of Toronto used a Custom CodeSet specifically designed to identify an Interferon signature in asymptomatic systemic autoimmune rheumatic diseases (SARDs) individuals. The ability to multiplex and the broad dynamic range enabled by the nCounter System allowed the researchers to identify and classify asymptomatic patients by their IFN signature, another step forward in the search for biomarkers of disease progression.
In another example regarding a transcriptional signature for multiple sclerosis (MS), Hu and colleagues from Brigham and Women’s Hospital at Harvard Medical School used a 418-gene Custom CodeSet specific for human T cell activation and differentiation to characterize a transcriptional signature of TH17 cells in MS. The team sorted rare CD4+ subsets —TH17 and TH1/TH17— and showed that the IL10 gene is reduced in MS compared to healthy controls. They found higher expression of the gene also in clinically stable vs. active patients, a finding that could define a molecular signature of human pro-inflammatory TH17 cells, which would, in turn, be used to measure the effect of treatment on TH17 cells in autoimmune disease.
Dr. Rogge Explains how Genetic Variants Affect Autoimmune Diseases
In his webinar, Dr. Lars Rogge, Director of the Immunoregulation Unit at the Institut Pasteur in Paris, shares with us his interest in closing the knowledge gap between the known genetic variants and underlying biological mechanisms of autoimmune disease. In a paper published in 2013, Dr. Rogge and his team demonstrated that the functions of both TH17 and TH1 cells in patients with Spondyloarthritis (SpA) are under combinatorial control by multiple SNPs at genes associated with the IL-23 signaling pathway, thus providing a framework to delineate the autoimmune pathways affected by genetic variants.
Following this evidence, the team decided to use the same approach to study how genetic variants affect signaling pathways involved in the pathogenesis of other chronic inflammatory diseases.
They encountered a challenge similar to what other researchers face when interrogating low-abundance immune cell samples, namely, how to find a robust and sensitive method to measure the expression of multiple genes from small cell populations.
As expected, NanoString nCounter technology was the answer. The team collaborated on the design of the nCounter® Autoimmune Discovery Panel, which is now available On-Demand. This panel allows for precise and robust expression analysis of 518 genes that have been found to be associated with risk variants identified in GWAS studies of nine chronic inflammatory diseases, as well as 237 genes involved in the regulation of immune responses. Watch the webinar to learn how the genes included in the panel were selected, curated and validated.
Watch these webinars On-Demand!
Take on the challenges of Autoimmunity research with the nCounter® Analysis System
References:
Joan Wither et al., Arthritis Res Ther. 2017; 19: 41.
Dan Hu et al, Nat Commun. 2017; 8: 1600.
Maryaline Coffre et al., ARTHRITIS & RHEUMATISM Vol. 65, No. 6, June 2013,
Read more on Autoimmunity:

Related Content